MSc. K. J. K. Lam, Dr. Z. Zhang, and Dr. M. Saier just published a paper in the Computational and Structural Biotechnology Journal. The title is: Histone-like nucleoid structuring (H-NS) protein silences the beta-glucoside (bgl) utilization operon in Escherichia coli by forming a DNA loop. For your convenience, this is the link to PubMed.
Abstract
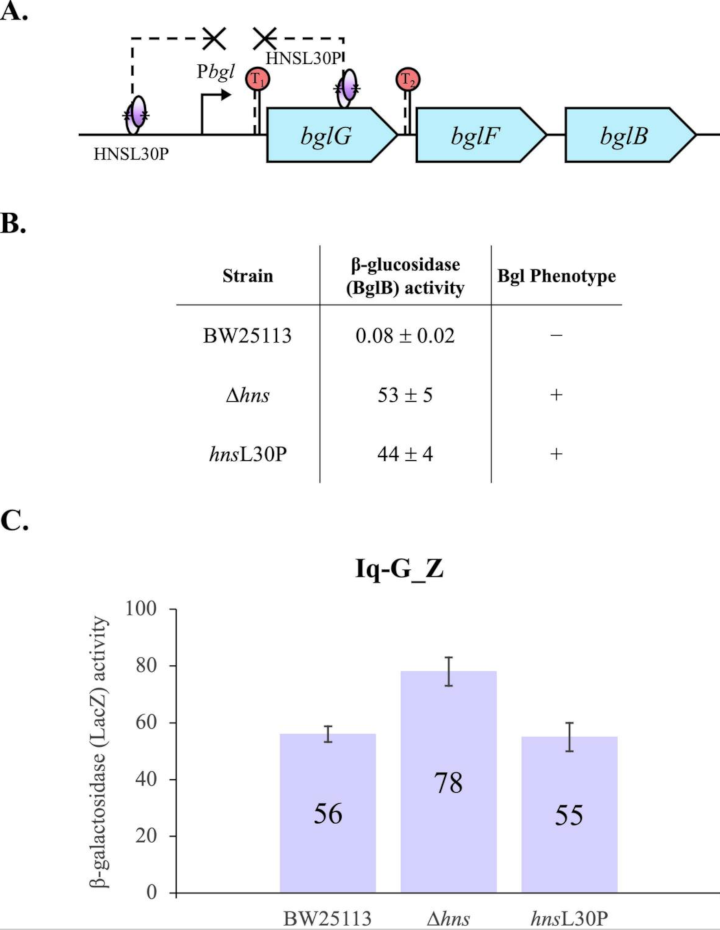
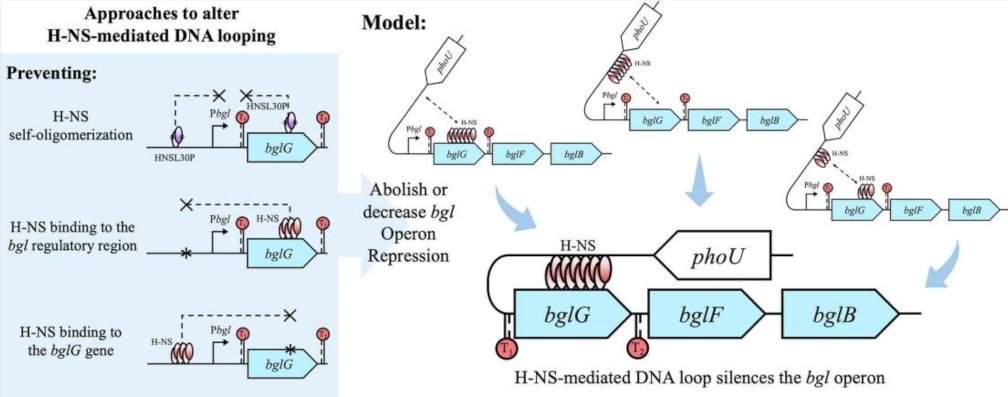
The bgl operon of Escherichia coli encodes proteins mediating the metabolism of aromatic beta-glucosides, but the operon is silent in wild type cells. Insertion of an insertion sequence (IS) element in the regulatory region upstream of the bgl promoter activates expression of the operon. The repression mechanism involves the histone-like nucleoid structuring (H-NS) protein with two DNA binding sites, one in the region upstream of the promoter, and the other within the first structural gene of the operon, bglG. The detailed mechanism of repression is not well understood. Here, we first show two terminators flanking bglG are not required for bgl operon silencing. Instead, several lines of experimental evidence clearly suggest that the silencing mechanism involves looping of the DNA between H-NS’s two DNA binding sites. H-NS is known to preferentially bind to AT-rich curved DNA, and such regions are found in the vicinity of both sites. We show that strong repression is abolished by (1) preventing H-NS self-oligomerization while retaining DNA binding, (2) preventing or reducing H-NS binding to the bgl operon regulatory region, and (3) preventing or reducing H-NS binding to the binding site in the bglG gene. We also show that the phase of the DNA between these two binding sites is not important, and that large insertions of DNA in the putative loop region do not diminish repression. These results imply that H-NS depends on DNA looping to exert strong repression.