Abstract:
Introduction: The transposase protein, InsAB’, is the key cytoplastic determinant that catalyzes IS1 movement within a single DNA molecule or between two DNA molecules. Regulation of InsAB’ synthesis, its structural characteristics, and its potent cis effect on IS1 transposition have been subjects of thorough investigation. However, limited efforts have been devoted to measuring the levels of IS1 transcripts and transposases made by native IS1 elements and examining their trans-acting function, particularly concerning their role in transposing otherwise immobile IS1 elements and other target DNA segments within the bacterial genome.
Methods: All constructs were made on the Escherichia coli chromosome using the lambda-Red system. IS1 transposition frequencies were determined using Bgl+ mutation assays, colony PCR and DNA sequencing. IS1 transcripts and transposases were quantitated using lacZ transcriptional and translational reporters.
Results: We first confirm that a native IS1 element, IS1E, can transpose at a dramatically elevated rate in the absence of both transcriptional and post-transcriptional regulations. Using lacZ reporters targeting the transposase gene insAB‘, we reveal that the InsA repressor moderately reduces insAB‘ transcription by as much as 16.8-fold. However, the combined action of InsA and ribosomal frameshifting leads to a much stronger reduction in InsAB’ production, by up to 735-fold. We show that only a small percentage (roughly 1.6%) of IS1 transcripts are successfully translated into InsAB’, highlighting the primary role of post-transcriptional regulation over transcriptional repression in governing IS1 transposition. We also demonstrate, for the first time, that high levels of InsAB’ exhibit a strong trans effect, being capable of efficiently transposing not only normally non-mobile IS1 elements but also a larger miniIS1 cassette within the genome, with greater amounts of InsAB’ leading to higher frequencies of transposition.
Conclusions: By dissecting and quantitating the roles of transcriptional and translational controls in transposase production, our study reveals that post-transcriptional regulation via ribosomal frameshifting is the central mechanism governing IS1 intragenomic transposition. Our findings demonstrate that IS1 transposases function in trans, mediating the movement of DNA segments specifically bordered by its inverted repeat at the left end and right end sequences. These findings advance our comprehension of how IS elements mobilize within bacterial genomes and provide potential strategies to counter IS1-mediated antibiotic resistance in clinical bacterial isolates.
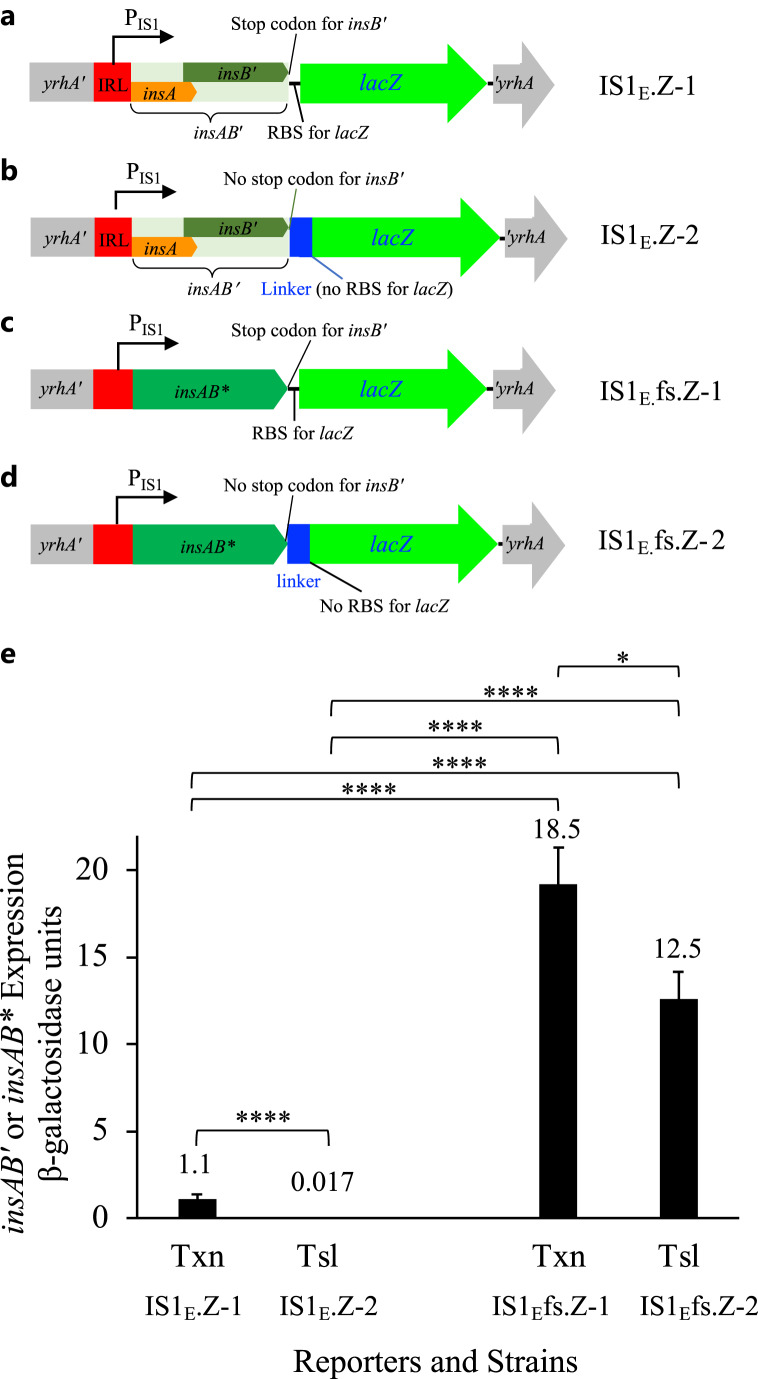
Figure 3. Quantifying IS1 transposase gene expression using lacZ reporters. a Diagram showing the lacZ transcriptional reporter for insAB′ within IS1E in strain IS1E.Z-1. insAB′ and lacZ, each with its own ribosome-binding site (RBS), form an operon that is driven by PIS1. b Diagram showing the lacZ translational reporter for insAB′ within IS1E in strain IS1E.Z-2. insAB′ and lacZ (without its own RBS), bridged by a 45-bp linker sequence, form a fusion gene that encodes a single complex protein via a ribosomal frameshift at the A6C motif. c Diagram showing the lacZ transcriptional reporter for insAB* within IS1Efs in strain IS1E.fs.Z-1. d Diagram showing the lacZ translational reporter for insAB* within IS1Efs in strain IS1E.fs.Z-2. insAB* and lacZ (without its own RBS) form a fusion gene that encodes a single complex protein without the ribosomal frameshift. e Quantitative measurements of transcriptional (n = 12 for each) and translational (n = 12 for each) levels of insAB′ within IS1E and insAB* within IS1Efs using β-galactosidase assays. “Txn” and “Tsl” denote transcriptional reporter and translational reporter, respectively. *p < 0.05 and ****p < 0.0001.


